King Tut’s DNA is Western European?
Source: eutimes.net

Despite the refusal of the Secretary General of the Egyptian Supreme Council of Antiquities, Zahi Hawass, to release any DNA results which might indicate the racial ancestry of Pharaoh Tutankhamen, the leaked results reveal that King Tut’s DNA is a 99.6 percent match with Western European Y chromosomes.
The DNA test results were inadvertently revealed on a Discovery Channel TV documentary filmed with Hawass’s permission — but it seems as if the Egyptian failed to spot the giveaway part of the documentary which revealed the test results.
Hawass previously announced that he would not release the racial DNA results of Egyptian mummies — obviously because he feared the consequences of such a revelation.
On the Discovery Channel broadcast, which can be seen on the Discovery Channel website here, or if they pull it, on YouTube here, at approximately 1:53 into the video, the camera pans over a printout of DNA test results from King Tut.
Firstly, here is a brief explanation of the results visible in the video. It is a list of what is called Short Tandem Repeats (STRs).
STRs are repeated DNA sequences which are “short repeat units” whose characteristics make them especially suitable for human identification.
These STR values for 17 markers visible in the video are as follows:
DYS 19 – 14 (? not clear)
DYS 385a – 11
DYS 385b – 14
DYS 389i – 13
DYS 389ii – 30
DYS 390 – 24
DYS 391 – 11
DYS 392 – 13
DYS 393 – 13
DYS 437 – 14 (? not clear)
DYS 438 – 12
DYS 439 – 10
DYS 448 – 19
DYS 456 – 15
DYS 458 – 16
DYS 635 – 23
YGATAH4 – 11
What does this mean? Fortunately, a genius by the name of Whit Athey provides the key to this list. Mr Athey is a retired physicist whose working career was primarily at the Food and Drug Administration where he was chief of one of the medical device labs.
Mr Athey received his doctorate in physics and biochemistry at Tufts University, and undergraduate (engineering) and masters (math) degrees at Auburn University. For several years during the 1980s, he also taught one course each semester in the electrical engineering department of the University of Maryland. Besides his interest in genetic genealogy, he is an amateur astronomer and has his own small observatory near his home in Brookeville, MD.
He also runs a very valuable website called the “Haplogroup Predictor” which allows users to input STR data and generate the haplogroup which marks those STR data.
For those who want to know what a haplogroup is, here is a “simple” definition: a haplogroup is a group of similar haplotypes that share a common ancestor with a single nucleotide polymorphism (SNP) mutation.
Still none the wiser? Damn these scientists.
Ok, let’s try it this way: a haplotype is a combination of multiple specific locations of a gene or DNA sequence on a chromosome.
Haplogroups are assigned letters of the alphabet, and refinements consist of additional number and letter combinations, for example R1b or R1b1. Y-chromosome and mitochondrial DNA haplogroups have different haplogroup designations. In essence, haplogroups give an inisight into ancestral origins dating back thousands of years.
By entering all the STR data inadvertently shown on the Discovery video, a 99.6 percent fit with the R1b haplogroup is revealed.
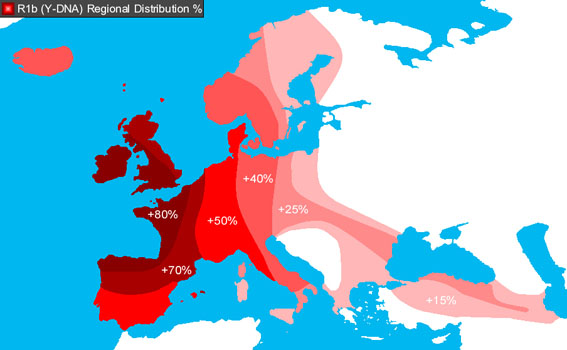
The significance is, of course, that R1b is the most common Y-chromosome haplogroup in Europe reaching its highest concentrations in Ireland, Scotland, western England and the European Atlantic seaboard — in other words, European through and through.
Article from: EUTimes.net






















